Early cost-effectiveness model of whole genome sequencing as a molecular diagnostic test versus standard of care in locally advanced and metastatic (Stage IIIB,C/ IV) non-squamous non-small cell lung cancer
Disease Area (Primary)
NSCLC
First Developed
07/25/2021
Last Developed
08/18/2021
Software Used
Microsoft Excel
Model License
Creative Commons (e.g., CC-BY, CC-BY-NC)
Model Sponsor
Government agency
Intervention
whole_genome_sequencing
– %
Coming Soon In Phase II: You will be able to pay a fee to download the CADTH Tool for your model which includes subaggregated scores.
Results
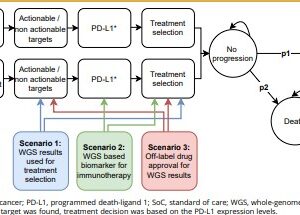
WGS as a diagnostic test resulted in more QALYs (0.002) and costs (€1534 [incremental net monetary benefit –€1349]), and SoC followed by WGS resulted in fewer QALYs (–0.002) and more costs (€1059 [–€1194]) compared with SoC alone. WGS as a diagnostic test was only cost effective if it was priced at €2000 per patient and identified 2.7% more actionable patients than SoC alone. Treating these additional identified patients with new treatments costing >€4069 per month decreased the probability of cost effectiveness.
Conclusion
Our analysis suggests that providing WGS as a diagnostic test would be cost effective compared with SoC followed by WGS and SoC alone if costs for WGS decreased and additional patients with actionable targets were identified. This cost-effectiveness model can be used to incorporate new findings iteratively and to support ongoing decision making regarding the use of WGS in this rapidly evolving field.
Source File(s)